What is aberrant splicing? Addressing the molecular foundations
Molecular biology is the scientific discipline committed to comprehend the diverse molecular structures and chemical processes that constitute the basis of cell interaction. Conceptually located right in between of genomics and biochemistry, this field has undergone game-changing transformation over the past decades, highlighting the discovery of DNA structure — Watson and Crick proposed their well-known double-helix conformation in a Nature publication back in 1953[1], setting the theoretical and experimental foundations for what was yet to come regarding further understanding of DNA nature and its molecular behaviour.
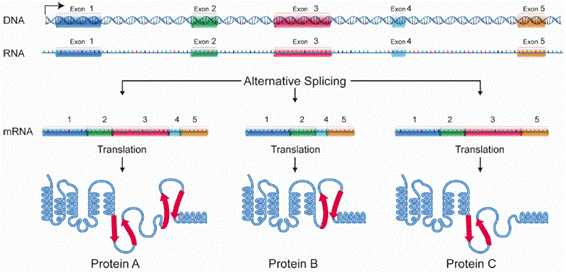
Among the countless number of processes that occur in cells at the molecular level, alternative splicing is one of a kind. First observed in 1977, it permits the obtention of different proteins — i.e., splice variants — from a single protein-coding gene[2]. That is possible because the minimal coding units of genes, known as exons, can either be skipped or included in the mature messenger RNA (mRNA) that will end up being converted to proteins in the cytoplasm. This alternance in the final structure of the mRNA, which eventually depends on both random and foreseeable factors, confers to alternative splicing its stochastic nature. Alternative splicing is a complex mechanism involving multiple steps until obtaining specific protein isoforms. It must be understood not only as a paramount biological process that contributes to protein diversity within the organism, but also as a natural source of diseases-causing errors in gene expression. For example, intronic splice site mutations of tumour suppressor genes often lead to exon-skipping events, thus truncating proteins similar to standard nonsense mutations[3]. Health problems stemming from this behaviour are often referred as aberrant alternative splicing (or simply aberrant splicing) events.
RNA Diseases — Wide spectrum of malignant outcomes
Transcription is the molecular process through which DNA is copied into messenger RNA (mRNA). Before being translated — that is, converted into proteins in the cytoplasm — there are multiple post-transcriptional processes that may take place, encompassing editing, modification, degradation, and splicing. We have addressed the latter, but it is important to understand that these defects or malfunctions can happen not only in splicing but also in any of the above-mentioned phenomena. Within this framework, human diseases caused by these events have been recently referred as “RNA diseases”[4], and we will follow this terminology.
RNA diseases compress neurological, muscular, cardiac, and cancer-related disorders. Some examples are included in the following table:

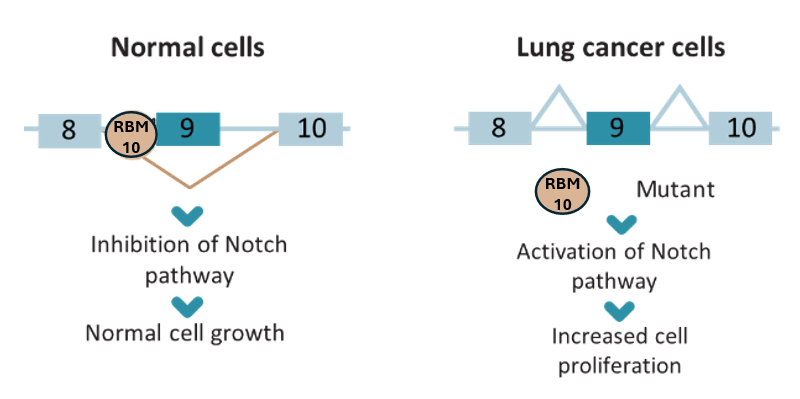
In literature, plenty of examples can be found regarding what are the exact genes whose misregulation triggers unwanted biochemical responses within cells, leading to some of the mentioned diseases. NUMB is an example of these genes and the main target for TAONas-LUAD, specifically its exon 9. Very briefly, NUMB gene encodes a protein — protein numb homolog — which is known to play a pivotal role in the determination of cell fates during development. In particular, it is key in asymmetrical cell division during neurogenesis (the process by which the nervous system creates new neurons), allowing for differential cell fate specification in the central and peripheral nervous systems[9]. Among the different exon that constitute NUMB, the one to watch for TAONas-LUAD researchers is exon 9. It has been shown how the inclusion/skipping or this exon in the mature mRNA is highly correlated with increased cell proliferation and consequent malignant outcomes in lung tissue[10], eventually leading to lung adenocarcinoma (i.e., LUAD).
And the list is not limited to NUMB: BRCA1 misregulation is associated with hereditary breast cancer, RB1 with retinoblastoma, KRAS with colorectal cancer, etc. Furthermore, cancer is only the tip of the iceberg, since many more disorders find their outset in gene misregulation.
TAONas-LUAD role — Committed to achieve clinical relevance
In TAONas-LUAD, this is our exact therapeutic focus:
“We try to disentangle the underlying molecular mechanisms behind aberrant splicing, and, with this know-how, we design tailor-made antisense oligonucleotides (AONs) aimed to revert the malignant biochemical cascades that lead to RNA diseases”
Designing AONs that effectively deliver therapeutic impact demands thorough understanding of the whole splicing process, as well as a well-thought, clinically-proven methodology encompassing optimal administration route, formulation, and dose. In TAONas-LUAD we have the privilege of working with many distinguished and experienced professionals in the field, ensuring that the correct steps are taken in this hard journey towards finding the right solution for aberrant splicing. The main focus of the project leaders as of now is placed is NUMB exon 9, but we slowly plan to broaden our scope and develop a full AON-based platform targeting aberrant splicing across numerous RNA diseases. Make sure you accompany us in this insightful adventure!
[1] Watson, J. D. & Crick, F. H. C. Molecular structure of nucleic acids: A structure for deoxyribose nucleic acid. Nature 171, 737–738 (1953).
[2] Chow, L. T., Gelinas, R. E., Broker, T. R. & Roberts, R. J. An amazing sequence arrangement at the 5′ ends of adenovirus 2 messenger RNA. Cell 12, 1–8 (1977).
[3] Venables, J. P. Aberrant and alternative splicing in cancer. Cancer Research 64, 7647–7654 (2004).
[4] Kataoka, N., Mayeda, A. & Ohno, K. Editorial: RNA Diseases in Humans—From Fundamental Research to Therapeutic Applications. Frontiers in Molecular Biosciences 6, 53 (2019).
[5] Nussbacher, J. K., Tabet, R., Yeo, G. W. & Lagier-Tourenne, C. Disruption of RNA metabolism in neurological diseases and emerging therapeutic interventions. Neuron 102, 294–320 (2019).
[6] Zavileyskiy, L. G. & Pervouchine, D. D. Post-transcriptional regulation of gene expression via unproductive splicing. Acta Naturae 16, 4–13 (2024).
[7] Shi, D.-L. RNA-Binding proteins in cardiomyopathies. Journal of Cardiovascular Development and Disease 11, 88 (2024).
[8] Carmo-Fonseca, M. & Valcárcel, J. The unfolding landscape of RNA and disease. RNA 31, 273–276 (2025).
[9] Pece, S., Confalonieri, S., Romano, P. R. & Di Fiore, P. P. NUMB-ing down cancer by more than just a NOTCH. Biochimica Et Biophysica Acta (BBA) – Reviews on Cancer 1815, 26–43 (2010).
[10] Bechara, E. G., Sebestyén, E., Bernardis, I., Eyras, E. & Valcárcel, J. RBM5, 6, and 10 differentially regulate NUMB alternative splicing to control cancer cell proliferation. Molecular Cell 52, 720–733 (2013).